This page was published for Genetics 564 at the University of Wisconsin-Madison
Protein interaction network for NSD1
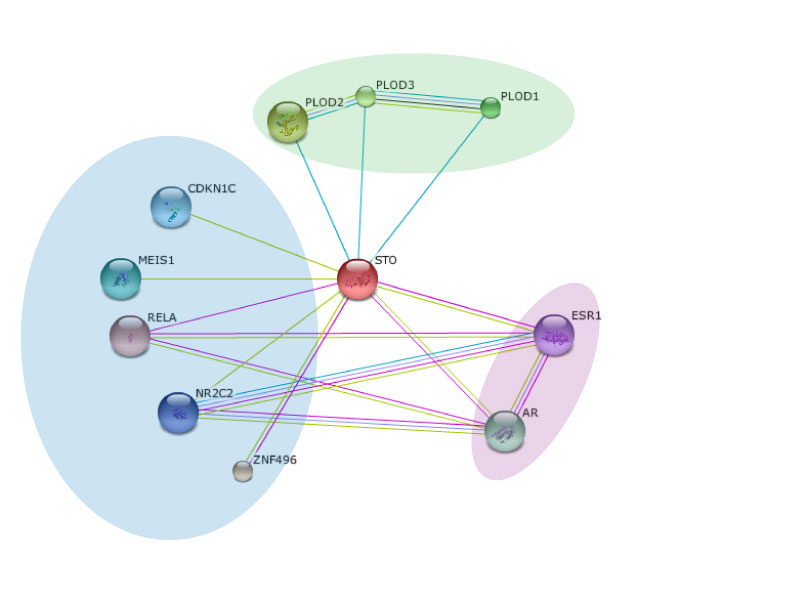
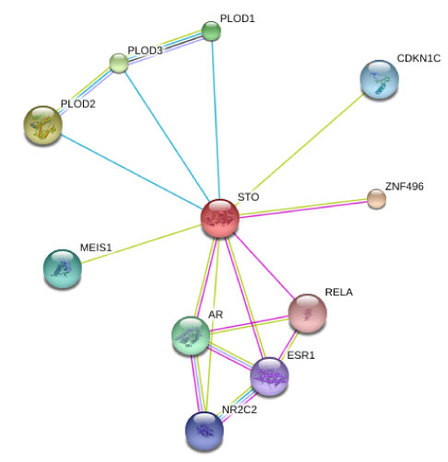 Figure 1: This is the protein interaction network for NSD1 (marked as STO)
Figure 1: This is the protein interaction network for NSD1 (marked as STO)
Protein-protein interactions are crucial for proper function of an organism. This occurs when at least two proteins associate to carry out a particular process or function. Fortunately, identifying interacting proteins has become easier than ever with the influx of genomic data available from the newly realized feasibility of sequencing entire genomes in a high-throughput manner. The difficult part comes with organization of this data [1]. This is where databases like STRING come into play, which allow the user to search for a particular protein of interest and obtain an interaction network, identifying key players with which the protein has been associated. The network corresponding to NSD1 is shown to the right (Figure 1). STRING uses data from researchers employing small scale protein-protein interaction assays such as TAP-tagging, Yeast Two Hybrid and Co-immunoprecipitation.
TAP tagging
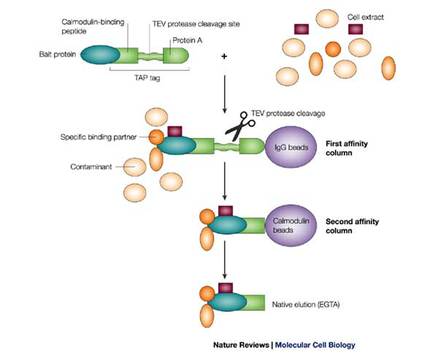 Figure 2: Schematic of the TAP tag assay
Figure 2: Schematic of the TAP tag assay
TAP tagging is an experiment that allows protein complex purification with the use of a small protein Tandem Affinity Purification (TAP) tag. This process begins by annealing a TAP tag to the protein of interest and introducing it to a host organism or cell under conditions where the protein will be expressed normally. Contents of the cell are then extracted and the protein of interest, now associated with its interacting partners, can be purified. The purification process is conducted in two steps. This is the unique feature of this method because it ensures minimal contamination.
The TAP tag has two notable IgG binding domains, the first called Protein A and the second called calmodulin binding protein (CBP) as can be seen in Figure 2 to the left. The two domains are separated by a protease cleavage site. The first step of the purification process involves the tight binding of Protein A to an IgG bead in solution. A protease is then introduced and the solution is washed to remove the IgG+Protein A complex along with any contaminants. The second step involves taking the remaining part of the protein-tag complex and exposing it to calmodulin coated beads in solution. The CBP domain of the tag then binds to these beads. This solution can be washed and rid of any extra lingering contaminants. The bound proteins can then be separated from the CBP+calmodulin bead under specific conditions. The result is a very pure mixture of the protein of interest and all of the associated proteins. These can be analyzed in a number of ways to determine the identity of interacting partners, structural characteristics, and other functional identifiers [2].
The TAP tag has two notable IgG binding domains, the first called Protein A and the second called calmodulin binding protein (CBP) as can be seen in Figure 2 to the left. The two domains are separated by a protease cleavage site. The first step of the purification process involves the tight binding of Protein A to an IgG bead in solution. A protease is then introduced and the solution is washed to remove the IgG+Protein A complex along with any contaminants. The second step involves taking the remaining part of the protein-tag complex and exposing it to calmodulin coated beads in solution. The CBP domain of the tag then binds to these beads. This solution can be washed and rid of any extra lingering contaminants. The bound proteins can then be separated from the CBP+calmodulin bead under specific conditions. The result is a very pure mixture of the protein of interest and all of the associated proteins. These can be analyzed in a number of ways to determine the identity of interacting partners, structural characteristics, and other functional identifiers [2].
Yeast Two Hybrid
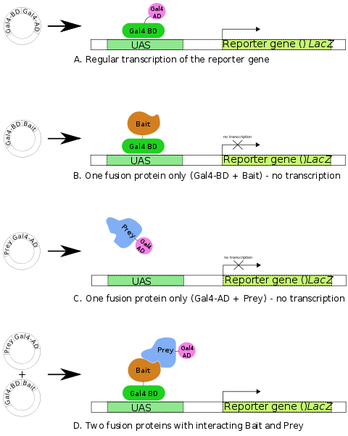 Figure 3: A schematic of the YTH assay depicting bait-prey interation
Figure 3: A schematic of the YTH assay depicting bait-prey interation
The Yeast Two Hybrid (YTH) system utilizes a yeast model to identify specific protein-protein interactions. In this system, a common transcription factor is broken into two parts: the DNA binding domain which gets fused to a bait protein and the activation domain of the transcriptional activator fused to a prey protein. As the bait and prey proteins physically interact, the transcription factor is once again active and the gene to which it corresponds is transcribed. This produces a measurable result [3]. This entire YTH process can be visualized in Figure 3 to the right. This assay is quite efficient and generally very easy to use, which makes it an attractive option compared to the TAP tag assay even though it is less accurate.
Co-immunoprecipitation
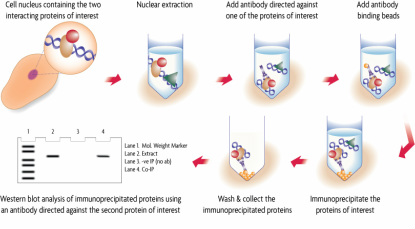 Figure 4: Schematic of the Co-IP assay
Figure 4: Schematic of the Co-IP assay
Another assay used to determine protein-protein interactions is called co-immunoprecipitation (Co-IP). This assay utilizes the inherent interaction of antibodies and antigens. Here, the bait protein is the known protein and acts as the antigen. The protein it interacts with is termed the prey. A cell containing these proteins of interest is lysed and then exposed to an antibody coupled with a bead. This antibody recognizes the bait and prey interaction formed by the cell and forms an immune complex. The solution is then washed and the immune complex isolated. The schematic of the Co-IP assay can be visualized in Figure 4 to the left. The bait and prey proteins remain bound throughout this assay, which can now be analyzed via Western blot or other procedure [4].
Results
I was able to retrieve the protein interaction network for NSD1 using the STRING database. The site allows manipulation of the image to stretch and move around the interacting partners like I did in the figure below. The different color lines connecting the proteins represent different interactions. Blue lines denote connections from various databases, purple lines denote experimental connections, and green lines show proteins in the same "neighborhood".
Analysis
After retrieving the protein interaction network from STRING for NSD1, I was able to categorize the interaction partners into a few groups. The three proteins circled in green belong to a class of proteins called PLOD, localized in the rough endoplasmic reticulum to catalyze the hydroxylation of lysines on collagen-like proteins. These hydroxyl groups then act as attachment sites for carbohydrates crucial for stable intermolecular linkages. The interaction between NSD1 and the PLOD proteins is not well characterized in Sotos syndrome, however the interaction may arise from the similar ability to modify lysine residues.
The two proteins circled in purple are steroid hormone receptors (estrogen and androgen). These interactions aren't specifically noted in any literature about Sotos syndrome either, however papers researching breast and prostate cancers have. This isn't too surprising given that patients who have Sotos tend to have a higher predisposition for tumor development [9]. Specifically, a paper researching the effectiveness of Tamoxifen as a drug in breast cancer in estrogen receptor-positive tumors found that the inhibition of NSD1 via RNAi corresponded to a resistance to the drug [5]. Furthermore, the histone methylation NSD1 carries out have been noted in carcinogenesis. High levels of histone methylation correspond with gene inactivation and in cancers, processes like tumor suppression and apoptosis are incapable of occurring. One paper noted this connection in prostate cancer (where androgen receptors are crucial) as a prognostic marker, specifically isolating the actions of NSD1's histone methyltransferase activities in recurring tumors [6]. NSD1 was shown to be down-regulated in the primary prostate tumors, however it was subsequently found to be upregulated in metastatic prostate tumors. The specific interactions between the androgen receptors in these tumors and NSD1 have not been clearly identified, but a correlation clearly exists. It appears that NSD1 has a role in providing an epigenetic signature for metastatic tumors that can potentially aid physicians in the diagnosis and characterization of recurring cancers [6].
The proteins circled in blue represent proteins involved with transcriptional regulation in development. Specifically cell proliferation and growth. CDKN1C has been noted as being mutated in Beckwith-Weidemann syndrome (BWS), which is another developmental disorder characterized by extreme overgrowth in childhood, much like Sotos syndrome. Although it is not the primary site of mutation in BWS, patients have been identified possessing mutations in this gene [7]. There is a high degree of clinical overlap between the two syndromes, and through a series of genetic analyses, a number of patients with Sotos were identified as having anomalies in the CDKN1C region. Conversely, a number of patients with BWS were identified as having anomalies in the NSD1 region [8]. Although the exact interaction between NSD1 and CDKN1C is unknown, it is thought that NSD1 works to imprint the chromosome on which CDKN1C resides [9].
MEIS1 is a growth-promoting oncogene associated with neuroblastoma, a type of cancer that develops in nerve tissue in infants and young children [10]. NSD1 function is known to be inhibited in neuroblastoma through transcriptional silencing via hypermethylation. This inhibition leads to the up-regulation of the MEIS1 oncogene, suggesting that MES1 is a target gene of NSD1 [11].
RELA is associated with cancer progression and the immune response and has been shown to interact with NSD1 through the Co-IP assay experimentally. NSD1 has been shown to drive the regulation pathway of this gene through its known histone methylation activity [12].
NR2C2 functions in spermatogenesis, cerebellum development, meiotic cell cycling and behavioral regulation [13]. NSD1 was shown to interact with NR2C2 through a YTH assay in the brain and the testes, suggesting interaction in these locations. Exact mechanism and result from this interaction is unclear [14].
ZNF496 is a zinc finger protein associated with transcriptional regulation. NSD1 is thought to bind directly to ZNF469 (also called Nizp1), which acts as a cofactor in transcriptional regulation. In Sotos mutations, however, this binding is unable to occur, and cognitive development is diminished- a notable characteristic in Sotos patients. Specifically, the PHD domains have been shown to bind the Nizp1 protein [15].
The two proteins circled in purple are steroid hormone receptors (estrogen and androgen). These interactions aren't specifically noted in any literature about Sotos syndrome either, however papers researching breast and prostate cancers have. This isn't too surprising given that patients who have Sotos tend to have a higher predisposition for tumor development [9]. Specifically, a paper researching the effectiveness of Tamoxifen as a drug in breast cancer in estrogen receptor-positive tumors found that the inhibition of NSD1 via RNAi corresponded to a resistance to the drug [5]. Furthermore, the histone methylation NSD1 carries out have been noted in carcinogenesis. High levels of histone methylation correspond with gene inactivation and in cancers, processes like tumor suppression and apoptosis are incapable of occurring. One paper noted this connection in prostate cancer (where androgen receptors are crucial) as a prognostic marker, specifically isolating the actions of NSD1's histone methyltransferase activities in recurring tumors [6]. NSD1 was shown to be down-regulated in the primary prostate tumors, however it was subsequently found to be upregulated in metastatic prostate tumors. The specific interactions between the androgen receptors in these tumors and NSD1 have not been clearly identified, but a correlation clearly exists. It appears that NSD1 has a role in providing an epigenetic signature for metastatic tumors that can potentially aid physicians in the diagnosis and characterization of recurring cancers [6].
The proteins circled in blue represent proteins involved with transcriptional regulation in development. Specifically cell proliferation and growth. CDKN1C has been noted as being mutated in Beckwith-Weidemann syndrome (BWS), which is another developmental disorder characterized by extreme overgrowth in childhood, much like Sotos syndrome. Although it is not the primary site of mutation in BWS, patients have been identified possessing mutations in this gene [7]. There is a high degree of clinical overlap between the two syndromes, and through a series of genetic analyses, a number of patients with Sotos were identified as having anomalies in the CDKN1C region. Conversely, a number of patients with BWS were identified as having anomalies in the NSD1 region [8]. Although the exact interaction between NSD1 and CDKN1C is unknown, it is thought that NSD1 works to imprint the chromosome on which CDKN1C resides [9].
MEIS1 is a growth-promoting oncogene associated with neuroblastoma, a type of cancer that develops in nerve tissue in infants and young children [10]. NSD1 function is known to be inhibited in neuroblastoma through transcriptional silencing via hypermethylation. This inhibition leads to the up-regulation of the MEIS1 oncogene, suggesting that MES1 is a target gene of NSD1 [11].
RELA is associated with cancer progression and the immune response and has been shown to interact with NSD1 through the Co-IP assay experimentally. NSD1 has been shown to drive the regulation pathway of this gene through its known histone methylation activity [12].
NR2C2 functions in spermatogenesis, cerebellum development, meiotic cell cycling and behavioral regulation [13]. NSD1 was shown to interact with NR2C2 through a YTH assay in the brain and the testes, suggesting interaction in these locations. Exact mechanism and result from this interaction is unclear [14].
ZNF496 is a zinc finger protein associated with transcriptional regulation. NSD1 is thought to bind directly to ZNF469 (also called Nizp1), which acts as a cofactor in transcriptional regulation. In Sotos mutations, however, this binding is unable to occur, and cognitive development is diminished- a notable characteristic in Sotos patients. Specifically, the PHD domains have been shown to bind the Nizp1 protein [15].
References:
[1] EMBL-EBI. 2014. The Importance of Molecular Interactions. Retrieved May 3, 2014 from http://www.ebi.ac.uk/training/online/course/protein-interactions-and-their-importance/protein-protein-interactions/importance-molecular-i
[2] Puig, O., Caspary, F., Rigaut, G., Rutz, B., Bouveret, E., Bragado-Nilsson, E., . . . Seraphin, B. (2001). The tandem affinity purification (TAP) method: a general procedure of protein complex purification. Methods, 24(3), 218-229. doi: 10.1006/meth.2001.1183
[3] Suter, B., Kittanakom, S., & Stagljar, I. (2008). Two-hybrid technologies in proteomics research. Current Opinion in Biotechnology, 19(4), 316-323. doi: http://dx.doi.org/10.1016/j.copbio.2008.06.005
[4] Sino Biological Inc. 2007. Co-immunoprecipitation Background and Protocol. Retrieved May 10, 2014 from http://www.assay-protocol.com/Immunology/Co-IP
[5] Mendes-Pereira, A. M., Sims, D., Dexter, T., Fenwick, K., Assiotis, I., Kozarewa, I., . . . Ashworth, A. (2012). Genome-wide functional screen identifies a compendium of genes affecting sensitivity to tamoxifen. Proc Natl Acad Sci U S A, 109(8), 2730-2735. doi: 10.1073/pnas.1018872108
[6] Bianco-Miotto, T., Chiam, K., Buchanan, G., Jindal, S., Day, T. K., Thomas, M., . . . Ricciardelli, C. (2010). Global levels of specific histone modifications and an epigenetic gene signature predict prostate cancer progression and development. Cancer Epidemiol Biomarkers Prev, 19(10), 2611-2622. doi: 10.1158/1055-9965.epi-10-0555
[7] Genetics Home Reference. 2014. Beckwith-Wiedemann syndrome. Retrieved May 16, 2014 from http://ghr.nlm.nih.gov/condition/beckwith-wiedemann-syndrome
[8] STRING. 2014. Textmining. Retrieved May 16, 2014 from http://string-db.org/newstring_cgi/show_textmining_evidence.pl?taskId=_Xw6jHHvsqMj&node1=987995&node2=997584
[9] Baujat, G., Rio, M., Rossignol, S., Sanlaville, D., Lyonnet, S., Le Merrer, M., . . . Colleaux, L. (2004). Paradoxical NSD1 mutations in Beckwith-Wiedemann syndrome and 11p15 anomalies in Sotos syndrome. Am J Hum Genet, 74(4), 715-720. doi: 10.1086/383093
[10] PubMed Health. 2013. Neuroblastoma. Retrieved May 16, 2014 from http://www.ncbi.nlm.nih.gov/pubmedhealth/PMH0002381/
[11] Berdasco, M., Ropero, S., Setien, F., Fraga, M. F., Lapunzina, P., Losson, R., . . . Esteller, M. (2009a). Epigenetic inactivation of the Sotos overgrowth syndrome gene histone methyltransferase NSD1 in human neuroblastoma and glioma. Proc Natl Acad Sci U S A, 106(51), 21830-21835. doi: 10.1073/pnas.0906831106
[12] Lu, T., Jackson, M. W., Wang, B., Yang, M., Chance, M. R., Miyagi, M., . . . Stark, G. R. (2010). Regulation of NF-kappaB by NSD1/FBXL11-dependent reversible lysine methylation of p65. Proc Natl Acad Sci U S A, 107(1), 46-51. doi: 10.1073/pnas.0912493107
[13] EMBL-EBI. 2014. QuickGO NR2C2 Homo Sapiens C9J2Y1. Retrieved May 16, 2014 from http://www.ebi.ac.uk/QuickGO/GProtein?ac=C9J2Y1
[14] STRING. 2014. Textmining. Retrieved May 16, 2014 from http://string-db.org/newstring_cgi/show_textmining_evidence.pl?taskId=_Xw6jHHvsqMj&node1=987995&node2=984596
[15] Pasillas, M. P., Shah, M., & Kamps, M. P. (2011). NSD1 PHD domains bind methylated H3K4 and H3K9 using interactions disrupted by point mutations in human sotos syndrome. Hum Mutat, 32(3), 292-298. doi: 10.1002/humu.21424
[1] EMBL-EBI. 2014. The Importance of Molecular Interactions. Retrieved May 3, 2014 from http://www.ebi.ac.uk/training/online/course/protein-interactions-and-their-importance/protein-protein-interactions/importance-molecular-i
[2] Puig, O., Caspary, F., Rigaut, G., Rutz, B., Bouveret, E., Bragado-Nilsson, E., . . . Seraphin, B. (2001). The tandem affinity purification (TAP) method: a general procedure of protein complex purification. Methods, 24(3), 218-229. doi: 10.1006/meth.2001.1183
[3] Suter, B., Kittanakom, S., & Stagljar, I. (2008). Two-hybrid technologies in proteomics research. Current Opinion in Biotechnology, 19(4), 316-323. doi: http://dx.doi.org/10.1016/j.copbio.2008.06.005
[4] Sino Biological Inc. 2007. Co-immunoprecipitation Background and Protocol. Retrieved May 10, 2014 from http://www.assay-protocol.com/Immunology/Co-IP
[5] Mendes-Pereira, A. M., Sims, D., Dexter, T., Fenwick, K., Assiotis, I., Kozarewa, I., . . . Ashworth, A. (2012). Genome-wide functional screen identifies a compendium of genes affecting sensitivity to tamoxifen. Proc Natl Acad Sci U S A, 109(8), 2730-2735. doi: 10.1073/pnas.1018872108
[6] Bianco-Miotto, T., Chiam, K., Buchanan, G., Jindal, S., Day, T. K., Thomas, M., . . . Ricciardelli, C. (2010). Global levels of specific histone modifications and an epigenetic gene signature predict prostate cancer progression and development. Cancer Epidemiol Biomarkers Prev, 19(10), 2611-2622. doi: 10.1158/1055-9965.epi-10-0555
[7] Genetics Home Reference. 2014. Beckwith-Wiedemann syndrome. Retrieved May 16, 2014 from http://ghr.nlm.nih.gov/condition/beckwith-wiedemann-syndrome
[8] STRING. 2014. Textmining. Retrieved May 16, 2014 from http://string-db.org/newstring_cgi/show_textmining_evidence.pl?taskId=_Xw6jHHvsqMj&node1=987995&node2=997584
[9] Baujat, G., Rio, M., Rossignol, S., Sanlaville, D., Lyonnet, S., Le Merrer, M., . . . Colleaux, L. (2004). Paradoxical NSD1 mutations in Beckwith-Wiedemann syndrome and 11p15 anomalies in Sotos syndrome. Am J Hum Genet, 74(4), 715-720. doi: 10.1086/383093
[10] PubMed Health. 2013. Neuroblastoma. Retrieved May 16, 2014 from http://www.ncbi.nlm.nih.gov/pubmedhealth/PMH0002381/
[11] Berdasco, M., Ropero, S., Setien, F., Fraga, M. F., Lapunzina, P., Losson, R., . . . Esteller, M. (2009a). Epigenetic inactivation of the Sotos overgrowth syndrome gene histone methyltransferase NSD1 in human neuroblastoma and glioma. Proc Natl Acad Sci U S A, 106(51), 21830-21835. doi: 10.1073/pnas.0906831106
[12] Lu, T., Jackson, M. W., Wang, B., Yang, M., Chance, M. R., Miyagi, M., . . . Stark, G. R. (2010). Regulation of NF-kappaB by NSD1/FBXL11-dependent reversible lysine methylation of p65. Proc Natl Acad Sci U S A, 107(1), 46-51. doi: 10.1073/pnas.0912493107
[13] EMBL-EBI. 2014. QuickGO NR2C2 Homo Sapiens C9J2Y1. Retrieved May 16, 2014 from http://www.ebi.ac.uk/QuickGO/GProtein?ac=C9J2Y1
[14] STRING. 2014. Textmining. Retrieved May 16, 2014 from http://string-db.org/newstring_cgi/show_textmining_evidence.pl?taskId=_Xw6jHHvsqMj&node1=987995&node2=984596
[15] Pasillas, M. P., Shah, M., & Kamps, M. P. (2011). NSD1 PHD domains bind methylated H3K4 and H3K9 using interactions disrupted by point mutations in human sotos syndrome. Hum Mutat, 32(3), 292-298. doi: 10.1002/humu.21424