This page was published for Genetics 564 at the University of Wisconsin- Madison
What is gene ontology?
The Gene Ontology (GO) project is the ongoing effort of classification of gene products across species. This bioinformatic tool has been produced for researchers searching to identify genes and gene products by offering a compilation of data from databases and labs across the world. In order to ensure functionally equivalent terminology- a key component of identification- the GO project has developed three vocabularies for universal description across all species. The three terms are listed below [1]:
GO terms:
GO terms:
- Molecular function: Molecular function refers to the activities occurring at the molecular level within the organism by individual gene products. This term encompasses the actions, not the actual proteins or molecules that perform the actions. Additionally, this term does not suggest the location or mechanism of the activities. Examples of molecular function GO terms include binding, receptor, or transporter activity.
- Biological processes: Biological processes are the actual events accomplished by one or more molecular functions. These two terms can be easily confused, but biological processes tend to include a series of steps, whereas molecular functions refer to a single activity by a gene product. Examples of biological process GO terms include signal transduction, metabolic processes, or photosynthetic processes.
- Cellular components: Cellular components are the locations of individual gene products within the cell. Examples of cellular components include the nucleus, endoplasmic reticulum, or cell membrane.
NSD1 GO data
To obtain this kind of information for NSD1, a search was done using AmiGO. The notable GO terms are listed below. For a full annotation of results, please click here. The flowcharts below were obtained from QuickGO.
Molecular functionChromatin binding
Histone methyltransferase activity Transcription corepressor activity Zinc ion binding Thyroid hormone receptor binding Estrogen receptor binding Androgen receptor binding |
Biological processesHistone methylation
Negative regulation of RNA polymerase II promoter Transcription, DNA templated Gastrulation with mouth forming second |
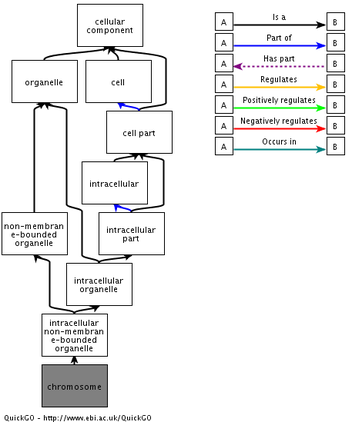
Cellular componentsChromosome
Nucleus |
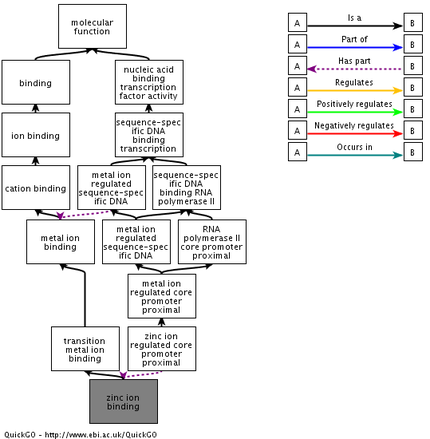
Flowcharts are useful tools to track specific GO terms in various pathways to see the scope of NSD1's function. Using the QuickGo database, three charts were created to outline GO terms from each of the tree categories. To the left is the flowchart corresponding to one of the molecular functions of NSD1, zinc ion binding. From this chart, it is easy to see how zinc ion binding contributes to transcription regulation. The literature has noted NSD1's regulatory behavior and further analysis of the domains show that there are a number of zinc finger binding domains on the protein, so these results make sense [2].
Below are the flowcharts exhibiting the ontology pathways of a novel cellular component and biological process associated with NSD1. Unsurprisingly, NSD1 has been localized amongst the chromosomes of the cell. This corresponds with the noted biological process of histone methylation, as histones are only found in chromosomes. The biological function chart has much higher levels of complexity as methylation of histones has been related to organization, metabolic processes, and protein alkylations.
Analysis
Time and time again, literature has noted the significance of histone methylation activity of the NSD1 protein [2][3][4]. In order for this process to be conducted properly, these three GO terms I have highlighted, plus a few more, must be in place. NSD1 is incapable of modifying histone structure without actually being located amongst the chromosomes inside the nucleus. Next, NSD1 must actually bind the chromatin structure through strong binding techniques such as zinc ion binding to produce the histone methyltransferase activity, that is, actually transferring the methyl groups to the appropriate histones. Lastly, the actual methylation of the histones will take place, directly affecting transcription through chromatin structure modification and RNA polymerase regulation [2].
There were a few extra molecular functions listed that did not directly relate to histone methylation, namely androgen and estrogen receptor binding. NSD1 has been liked to other proteins associated with these processes, but information directly related to Sotos syndrome does not primarily highlight these activities. Further research could be conducted to determine fertility of patients with Sotos or the development of secondary sex characteristics during puberty.
There were a few extra molecular functions listed that did not directly relate to histone methylation, namely androgen and estrogen receptor binding. NSD1 has been liked to other proteins associated with these processes, but information directly related to Sotos syndrome does not primarily highlight these activities. Further research could be conducted to determine fertility of patients with Sotos or the development of secondary sex characteristics during puberty.
References:
[1] The Gene Ontology Consortium. Gene ontology: tool for the unification of biology. Nat. Genet. May 2000; 25(1): 25-9.
[2] Qiao, Q., Li, Y., Chen, Z., Wang, M., Reinberg, D., & Xu, R. M. (2011). The structure of NSD1 reveals an autoregulatory mechanism underlying histone H3K36 methylation. J Biol Chem, 286(10), 8361-8368. doi: 10.1074/jbc.M110.204115
[3] Carpenter, P. B. (2014). Description of Research. Epigenetics and the NSD family in development and disease. Retrieved March 5, 2014, 2014
[4] Faravelli, F. (2005). NSD1 mutations in Sotos syndrome. American Journal of Medical Genetics, 137C(1), 24-31. doi: 10.1002/ajmg.c.30061
[1] The Gene Ontology Consortium. Gene ontology: tool for the unification of biology. Nat. Genet. May 2000; 25(1): 25-9.
[2] Qiao, Q., Li, Y., Chen, Z., Wang, M., Reinberg, D., & Xu, R. M. (2011). The structure of NSD1 reveals an autoregulatory mechanism underlying histone H3K36 methylation. J Biol Chem, 286(10), 8361-8368. doi: 10.1074/jbc.M110.204115
[3] Carpenter, P. B. (2014). Description of Research. Epigenetics and the NSD family in development and disease. Retrieved March 5, 2014, 2014
[4] Faravelli, F. (2005). NSD1 mutations in Sotos syndrome. American Journal of Medical Genetics, 137C(1), 24-31. doi: 10.1002/ajmg.c.30061