This page was published for Genetics 564 at the University of Wisconsin-Madison
Conclusions
Sotos syndrome is a highly complex disease associated with a plethora of symptoms. Although many of the most notable aspects of Sotos fade with age (facial gestalt, abnormal height, large head), there are a few that remain throughout the lifetime. One of these include serious bone ailments resulting from the advanced bone age, such as scoliosis. Another, which may be the most significantly lasting characteristic of Sotos syndrome, is a learning impairment. Patients can have learning disabilities ranging in severity from mild to extreme. Some adults with Sotos are able to live on their own, hold a job and even have a family, whereas others require constant assistance and are unable to care for themselves entirely [1]. There is a significant lack of research surrounding these learning disabilities, specifically noting what is responsible for the range in severity and what the underlying mechanism is. Throughout my research, I developed a few future directions that could be taken in to help better characterize this part of Sotos syndrome.
Although it would be ideal to use humans or chimpanzees as a research model, I kept in mind cost and feasibility when designing the following experiments. This is why I chose to use a Drosophila melanogaster (fruit fly) model. Fruit flies posses a homolog of NSD1 called Mes-4 that shares 36% identity with humans, possesses almost all of the notable domains of NSD1 and is easily genetically manipulated. One of these domains, the SET domain appears to dictate the function of the protein. This is the domain that carries out the actual function of the histone methylation, and I believe that mutations in this domain could be what is causing the disabilities in the first place.
I arrived at this hypothesis because histone methylation has been connected to learning and memory in past studies. One group of researchers used a mouse model to characterize this connection by sending mice through a conditioned learning assay assessing the ability to associate a sound with a foot shock. The mice were tested and their hippocampi extracted to analyze the levels of histone methylation at the H3 site on the histones, which, incidentally, is the same location where NSD1 methylates. Researchers then knocked out a gene responsible for histone methylation (not NSD1) and conducted the same conditioning assay. These mice were unable to show the same learned response and, unsurprisingly, had decreased levels of methylation of their histones [2].
I took the information learned from this study and identified a fairly apparent jumping off point for analysis of NSD1. What is the function of NSD1 in learning, exactly? To answer this, I developed the following experiments.
Although it would be ideal to use humans or chimpanzees as a research model, I kept in mind cost and feasibility when designing the following experiments. This is why I chose to use a Drosophila melanogaster (fruit fly) model. Fruit flies posses a homolog of NSD1 called Mes-4 that shares 36% identity with humans, possesses almost all of the notable domains of NSD1 and is easily genetically manipulated. One of these domains, the SET domain appears to dictate the function of the protein. This is the domain that carries out the actual function of the histone methylation, and I believe that mutations in this domain could be what is causing the disabilities in the first place.
I arrived at this hypothesis because histone methylation has been connected to learning and memory in past studies. One group of researchers used a mouse model to characterize this connection by sending mice through a conditioned learning assay assessing the ability to associate a sound with a foot shock. The mice were tested and their hippocampi extracted to analyze the levels of histone methylation at the H3 site on the histones, which, incidentally, is the same location where NSD1 methylates. Researchers then knocked out a gene responsible for histone methylation (not NSD1) and conducted the same conditioning assay. These mice were unable to show the same learned response and, unsurprisingly, had decreased levels of methylation of their histones [2].
I took the information learned from this study and identified a fairly apparent jumping off point for analysis of NSD1. What is the function of NSD1 in learning, exactly? To answer this, I developed the following experiments.
Experiment 1
To determine which domains in NSD1/Mes-4 contribute to learning
Approach: To conduct this experiment, I plan on utilizing a structure function analysis on Mes-4 in Drosophila and running the mutant flies through a conditioning learning assay. The structure function analysis will allow differentiation between the function of each specific domain in learning.
Hypothesis: I hypothesize that a mutation in the SET domain (i.e., removing the SET domain entirely) will cause the most profound effect in learning.
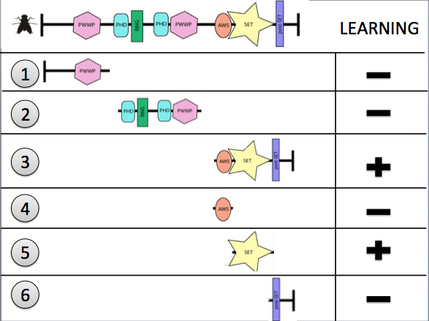
Discussion: Once the mutant flies are made as depicted in Figure 1 below, the flies will be run through an olfactory learning assay. This assay takes each mutant fly, places them in a tube with one odor and the administration of a series of slight shocks. The flies are then placed in a second tube and exposed to a second odor, except this time without shocks. After training, the flies will be entered into a T-maze as shown in Figure 2. One end of the T-maze will emit odor 1 (associated with shock) and the other end will emit odor 2 (no shock). The flies will be allowed to localize at either end of the maze, counted and compared to the control to assess learning ability. I predict that out of the 6 different potential structures of Mes-4, the mutants without the SET domain would have a decreased learning response compared to those with the domain included.
Approach: To conduct this experiment, I plan on utilizing a structure function analysis on Mes-4 in Drosophila and running the mutant flies through a conditioning learning assay. The structure function analysis will allow differentiation between the function of each specific domain in learning.
Hypothesis: I hypothesize that a mutation in the SET domain (i.e., removing the SET domain entirely) will cause the most profound effect in learning.
Discussion: Once the mutant flies are made as depicted in Figure 1 below, the flies will be run through an olfactory learning assay. This assay takes each mutant fly, places them in a tube with one odor and the administration of a series of slight shocks. The flies are then placed in a second tube and exposed to a second odor, except this time without shocks. After training, the flies will be entered into a T-maze as shown in Figure 2. One end of the T-maze will emit odor 1 (associated with shock) and the other end will emit odor 2 (no shock). The flies will be allowed to localize at either end of the maze, counted and compared to the control to assess learning ability. I predict that out of the 6 different potential structures of Mes-4, the mutants without the SET domain would have a decreased learning response compared to those with the domain included.
Experiment 2
To determine which genes and proteins are necessary for learning
Approach: For this experiment I plan on utilizing two separate sub-approaches. The first sub-approach is to use STRING to identify binding partners for which NSD1 interacts and organize the results by Gene Ontology (GO) to identify which are associated with learning. Then, take these results and run them through SMART to identify any crucial domains and compare the results to NSD1's domain profile. The second sub-approach is utilizing RNA-seq in the Drosophila model and then organizing the expression profile by GO terms for learning. The results will be compared between wild type and SET mutants to observe which terms are different.
Hypothesis: I predict that there will be a wide variety of different proteins and genes associated with learning and many have SET domains.
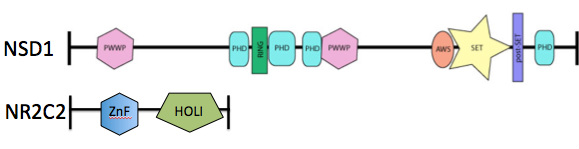
Discussion: I actually conducted the first sub-approach and from the STRING analysis, I found that of all the interacting partners, there was one protein (NR2C2) that listed "behavior regulation" under the GO terms. I ran NR2C2 through SMART and discovered that there were no SET domains at all. In fact, there were no similar domains in NSD1 and NR2C2 whatsoever. The two domains seen in NR2C2 are known to function in DNA binding. And, from all that is known on the domains in NSD1, some of these function in DNA binding as well. This leads to the conclusion that SET domains may not be crucial for learning, whereas DNA binding-associated domains are. The two proteins are seen aligned in Figure 3.
The second sub-approach, I predict, will result in two sets of gene expression profiles. The first associated with the wild type flies and the second associated with the SET mutants, that is, it is missing the SET domain. I hypothesize that there will be a subset of learning-related GO terms that exist in the WT flies, but are absent in the SET mutants. In order to confirm that these genes are indeed related to learning, I will knock them out in flies and run them through the learning assay discussed in Experiment 1. Then I will assess the learning ability.
Approach: For this experiment I plan on utilizing two separate sub-approaches. The first sub-approach is to use STRING to identify binding partners for which NSD1 interacts and organize the results by Gene Ontology (GO) to identify which are associated with learning. Then, take these results and run them through SMART to identify any crucial domains and compare the results to NSD1's domain profile. The second sub-approach is utilizing RNA-seq in the Drosophila model and then organizing the expression profile by GO terms for learning. The results will be compared between wild type and SET mutants to observe which terms are different.
Hypothesis: I predict that there will be a wide variety of different proteins and genes associated with learning and many have SET domains.
Discussion: I actually conducted the first sub-approach and from the STRING analysis, I found that of all the interacting partners, there was one protein (NR2C2) that listed "behavior regulation" under the GO terms. I ran NR2C2 through SMART and discovered that there were no SET domains at all. In fact, there were no similar domains in NSD1 and NR2C2 whatsoever. The two domains seen in NR2C2 are known to function in DNA binding. And, from all that is known on the domains in NSD1, some of these function in DNA binding as well. This leads to the conclusion that SET domains may not be crucial for learning, whereas DNA binding-associated domains are. The two proteins are seen aligned in Figure 3.
The second sub-approach, I predict, will result in two sets of gene expression profiles. The first associated with the wild type flies and the second associated with the SET mutants, that is, it is missing the SET domain. I hypothesize that there will be a subset of learning-related GO terms that exist in the WT flies, but are absent in the SET mutants. In order to confirm that these genes are indeed related to learning, I will knock them out in flies and run them through the learning assay discussed in Experiment 1. Then I will assess the learning ability.
Experiment 3
To identify conserved amino acids important for learning in mammals
Approach: For this experiment, I plan on using Clustal Omega to align all of the primary sequences for the protein homologs of humans, chimpanzee, mouse, fruit fly, yeast and Arabidopsis.
Hypothesis: I hypothesize that mammals will have a different set of amino acids in the SET domain than non-mammals.
Discussion: After running these alignments through Clustal Omega, I found that the area of most significant overlap actually appeared in a non-domain region at the C-terminus end of the protein (as can be seen in Figure 4). In this location, there are a number of sites that can be seen to have significant differences in the mammals and non-mammals. I also noticed a trend where the fruit flies did not identify with the mammal group or yeast + arabidopsis group. I believe this could be because although mammals and flies are capable of learning, flies are not able to learn at the same intellectual level as the mammals, however they are still able to learn better than yeast + arabidopsis.
Hypothesis: I hypothesize that mammals will have a different set of amino acids in the SET domain than non-mammals.
Discussion: After running these alignments through Clustal Omega, I found that the area of most significant overlap actually appeared in a non-domain region at the C-terminus end of the protein (as can be seen in Figure 4). In this location, there are a number of sites that can be seen to have significant differences in the mammals and non-mammals. I also noticed a trend where the fruit flies did not identify with the mammal group or yeast + arabidopsis group. I believe this could be because although mammals and flies are capable of learning, flies are not able to learn at the same intellectual level as the mammals, however they are still able to learn better than yeast + arabidopsis.
Although there are a few different locations of mismatch, a few notable sites are highlighted in Figure 5. Here, it can be seen that the amino acids in mammals (threonine or serine) are phosphorylated whereas the amino acids in yeast + arabidopsis (alanine or methionine) are acetylated. Fruit flies, on the other hand, had inconsistencies. As previously mentioned, flies are not consistent with the patterns of mammal group or the yeast + arabidopsis group. Furthermore, it has been noted that acetylations and phosphorylations have an effect on transcription, so it could be interesting to see how, say, a mammal who normally has an amino acid that was phosphorylated would react if that amino acid was changed to one that could be acetylated and note the difference in learning.
Future directions
Analyze non-domain regions
I'd like to further analyze non-domain regions, because as specified from Experiment 3, there is some significant differences to be seen between learning-capable organisms and learning-incapable organisms in these locations. A structure function analysis assay could be used here and run through a learning assay much like in Experiment 1.
Classify learning disability types
There are over 100 different mutations in NSD1 that are associated with Sotos syndrome. I believe it could be interesting to classify the different mutations with different kinds of learning disabilities. There are many different kinds of learning, including visual learning, associative learning, auditory learning, etc. So I think it could be interesting to classify these differences in accordance to the type of mutation each patient has and determine if there is any correlation.
Determine learning disabilities in males and females
NSD1 has been shown to interact with androgen receptors (AR) and estrogen receptors (ER). I think it could be interesting to see how different learning disabilities manifest in male and female Sotos patients. This could provide further interaction information between NSD1 and these proteins and help to better characterize the exact mechanism of learning capabilities in these patients.
I'd like to further analyze non-domain regions, because as specified from Experiment 3, there is some significant differences to be seen between learning-capable organisms and learning-incapable organisms in these locations. A structure function analysis assay could be used here and run through a learning assay much like in Experiment 1.
Classify learning disability types
There are over 100 different mutations in NSD1 that are associated with Sotos syndrome. I believe it could be interesting to classify the different mutations with different kinds of learning disabilities. There are many different kinds of learning, including visual learning, associative learning, auditory learning, etc. So I think it could be interesting to classify these differences in accordance to the type of mutation each patient has and determine if there is any correlation.
Determine learning disabilities in males and females
NSD1 has been shown to interact with androgen receptors (AR) and estrogen receptors (ER). I think it could be interesting to see how different learning disabilities manifest in male and female Sotos patients. This could provide further interaction information between NSD1 and these proteins and help to better characterize the exact mechanism of learning capabilities in these patients.
| masliahjamiepresentation.pdf | |
| File Size: | 6574 kb |
| File Type: | |
References:
[1] Cole, TRP., Hughes, HE. (1994). Sotos syndrome: a study of the diagnostic criteria and natural history. Journal of Medical Genetics, 31. 20-32. doi: 10.1136/jmg.31.1.20
[2] Gupta, S., Kim, S. Y., et.al. (2010). Histone methylation regulates memory formation. The Journal of Neuroscience. 30(10). 3589-3599.
[1] Cole, TRP., Hughes, HE. (1994). Sotos syndrome: a study of the diagnostic criteria and natural history. Journal of Medical Genetics, 31. 20-32. doi: 10.1136/jmg.31.1.20
[2] Gupta, S., Kim, S. Y., et.al. (2010). Histone methylation regulates memory formation. The Journal of Neuroscience. 30(10). 3589-3599.