This page was published for Genetics 564 at the University of Wisconsin-Madison
What are domains and motifs?
Domains are the functional units of proteins. They are conserved folded segments that have a specific function or interaction pattern that dictates the overall role of the protein. Additionally, domains possess the ability to function independently. Motifs, on the other hand, are highly conserved amino acid sequences that make up the primary structure of the protein [1]. These domains and motifs work together to form the functionally relevant protein needed by the cell.
Proteins can be comprised of one or many domains. Multi-domain proteins, like NSD1, exhibit a number of interactions between domains that work together to direct the protein's function. These types of proteins have been thought to arise from duplications and recombinations within genes, resulting in a multi-domain protein with new or altered functions [2].
Proteins can be comprised of one or many domains. Multi-domain proteins, like NSD1, exhibit a number of interactions between domains that work together to direct the protein's function. These types of proteins have been thought to arise from duplications and recombinations within genes, resulting in a multi-domain protein with new or altered functions [2].
Protein domains of human NSD1 isoform b

Figure 1: This figure denotes the key protein domains found in human NSD1 using PFAM. The key domains include 2 PWWP domains, a PHD- finger domain, and a SET domain.
Figure 2a: This figure denotes the key protein domains found in human NSD1 using SMART. The noted domains include 2 PWWPs, 4 PHDs, SET and postSET, RING, and AWS.
Domains in the NSD1 protein are plentiful and important. The key players noted by numerous papers and the SMART results listed above are the PWWP, PHD, RING, and SET (with pre/postSET) domains. These specific regions interact with one another to form the functional histone methyltransferase activity of the protein. Although the exact function of NSD1 isn't entirely known, the functions of each domain can help to elucidate how it functions on the chromosomal level in transcriptional regulation.
Different forms of proteins can arise from the same gene via alternative splicing. This mechanism acts to create a myriad of gene products called isoforms that commonly have a variety of functions [3]. There is one other noted isoform of the NSD1 protein, NSD1 isoform a (Figure 2b). This alternate form of the NSD1 protein has a very similar structure as determined by the domains. Isoform a includes the noted SET and postSET, PWWP, and PHD domains but lacks a RING and substitutes a different preSET domain. There hasn't been much insight into the difference in function between the two isoforms, but one can deduce the high probability of similar function from comparison of the two domain structures.
Different forms of proteins can arise from the same gene via alternative splicing. This mechanism acts to create a myriad of gene products called isoforms that commonly have a variety of functions [3]. There is one other noted isoform of the NSD1 protein, NSD1 isoform a (Figure 2b). This alternate form of the NSD1 protein has a very similar structure as determined by the domains. Isoform a includes the noted SET and postSET, PWWP, and PHD domains but lacks a RING and substitutes a different preSET domain. There hasn't been much insight into the difference in function between the two isoforms, but one can deduce the high probability of similar function from comparison of the two domain structures.
Figure 2b: This figure denotes the key protein domains in human NSD1 isoform a using SMART. The noted domains include 2 PWWPs, 5 PHDs, SET and postSET, and a preSET EFG-like site.
PWWP domain
The proline-tryptophan-tryptophan-proline domain or PWWP, as denoted by the single amino acid letter sequence, is known to bind to the methylated lysine-20 on histone 4 (H4K20me) [3]. The close proximity to other domains suggests that PWWP has some role in protein-protein interactions [4]. PWWP's histone methylation capabilities correspond to a significant role in the organization of higher-order chromatin, maintenance of genome stabilization and the regulation of cell-cycle progression [3].

Figure 3: This is the crystalline structure of the PWWP domain notably found in the human NSD1 protein [3].
PHD domain
The plant homeodomain (PHD) finger is a zinc finger-like domain commonly associated with proteins involved in chromatin-based transcriptional regulation. Structurally similar to the RING domain, the PHD binds two zinc ions with finger-like protrusions to aid in molecular function [5]. Reportedly, PHD binds to trimethylated lysine-4 on histone 3 (H3K4me3) and acts as an epigenetic reader seeing as though it binds to the modified form of the histone. These interactions allow dynamic modifications to occur [6].
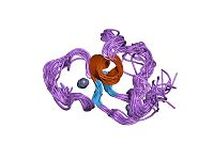
Figure 4: This is the crystalline structure of the PHD domain notably found in human NSD1 protein. The zinc ions are shown in gray [6].
SET and postSET domains
The SET domain was originally identified in Drosophila melanogaster Su(var)3-9, "Enhancer of zeste", and Trithorax proteins where researchers obtained the acronym SET. Commonly associated in a string of protein domains, SET acts with pre- and post-SET domains to take part in chromosomal modifications, much like the other domains listed here. It has been noted that in the C-terminal motif of the domain, tyrosine is highly conserved and acts to grab a proton off of the protonated amino group of lysine on the histones (most commonly H3). This then facilitates the nucleophilic attack on the AdoMet cofactor (a small protein that aids in methyl group transfers) to result in the addition/subtraction of one methyl group from the target histone. This directly relates to the notion of chromosomal modification [7].
The postSET domain comes directly after the SET domain, as the name suggests. It is located at the C terminal of the SET domain and is made of a series of highly conserved cysteine residues that form a zinc binding site. This brings in the required residues for histone tail interactions.
The postSET domain comes directly after the SET domain, as the name suggests. It is located at the C terminal of the SET domain and is made of a series of highly conserved cysteine residues that form a zinc binding site. This brings in the required residues for histone tail interactions.
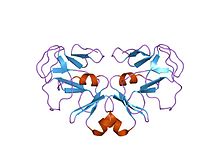
Figure 5: This is the crystalline structure of the SET domain [7].
RING domain
The RING domain is a zinc finger that functions through tandem binding of target molecules to mediate protein-protein interactions. RING is involved with a variety of biological processes ranging from transcription and translation to cell adhesion and chromatin remodeling [8].
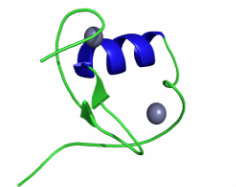
Figure 6: This is the crystalline structure of the RING domain [8].
Domain homology between model organisms for NSD1
Figure 7: This figure depicts the homology between model organisms containing homologous proteins to NSD1. For more information concerning the various protein homologs, please visit the Protein Homology page.
Analysis
Although the PFAM and SMART protein domain results showed different structures of importance for NSD1, the literature remains constant in identifying the PWWP, PHD, RING, pre/postSET and SET domains as significant. Homology throughout model organisms also points to importance in these defined domains. The histone methyltransferase activity is well conserved in these organisms, as are the pre/postSET and SET domains, suggesting that this is the crucial segment of the NSD1 protein required for this type of transcriptional regulation at the specific histone sites.
References:
[1] Protein Structure and Structural Bioinformatics. (2014). Basics of Protein Structure. Retrieved April 23, 2014 from http://www.proteinstructures.com/Structure/Structure/protein-domains.html
[2] Apic, G., Gough, J., & Teichmann, S. A. (2001). Domain combinations in archaeal, eubacterial and eukaryotic proteomes. Journal of Molecular Biology, 310(2), 311-325. doi: http://dx.doi.org/10.1006/jmbi.2001.4776
[3] PFAM. (2014). Summary: PWWP domain. Retrieved March 4, 2014 from http://pfam.sanger.ac.uk/family/PF00855.12
[4] SMART. (2014). PWWP. Retrieved March 4, 2014 from http://smart.embl-heidelberg.de/smart/do_annotation.pl?DOMAIN=PWWP&START=321&END=387&E_VALUE=0.00102386718599062&TYPE=SMART&BLAST=YEVGDLIWAKFKRRPWWPCRICSDPLINTHSKMKVSNRRPYRQYYVEAFGDPSERAWVAGKAIVMFE
[5] SMART. (2014). PHS. Retrieved March 5, 2014 from http://smart.embl-heidelberg.de/smart/do_annotation.pl?DOMAIN=PHD&START=1640&END=1693&E_VALUE=10.8700080063499&TYPE=SMART&BLAST=CITCHAANPANVSASKGRLMRCVRCPVAYHANDFCLAAGSKILASNSIICPNHF
[6] PFAM. (2014). Summary: PHD finger. Retrieved March 5, 2014 from http://pfam.sanger.ac.uk/family/PF00628.24
[7] PFAM. (2014). Summary: SET domain. Retrieved March 5, 2014 from http://pfam.sanger.ac.uk/family/PF00856.23
[8] The Pawson Lab. (2014). RING domain. Retrieved March 6, 2014 from http://pawsonlab.mshri.on.ca/index.php?option=com_content&task=view&Itemid=64&id=176
[1] Protein Structure and Structural Bioinformatics. (2014). Basics of Protein Structure. Retrieved April 23, 2014 from http://www.proteinstructures.com/Structure/Structure/protein-domains.html
[2] Apic, G., Gough, J., & Teichmann, S. A. (2001). Domain combinations in archaeal, eubacterial and eukaryotic proteomes. Journal of Molecular Biology, 310(2), 311-325. doi: http://dx.doi.org/10.1006/jmbi.2001.4776
[3] PFAM. (2014). Summary: PWWP domain. Retrieved March 4, 2014 from http://pfam.sanger.ac.uk/family/PF00855.12
[4] SMART. (2014). PWWP. Retrieved March 4, 2014 from http://smart.embl-heidelberg.de/smart/do_annotation.pl?DOMAIN=PWWP&START=321&END=387&E_VALUE=0.00102386718599062&TYPE=SMART&BLAST=YEVGDLIWAKFKRRPWWPCRICSDPLINTHSKMKVSNRRPYRQYYVEAFGDPSERAWVAGKAIVMFE
[5] SMART. (2014). PHS. Retrieved March 5, 2014 from http://smart.embl-heidelberg.de/smart/do_annotation.pl?DOMAIN=PHD&START=1640&END=1693&E_VALUE=10.8700080063499&TYPE=SMART&BLAST=CITCHAANPANVSASKGRLMRCVRCPVAYHANDFCLAAGSKILASNSIICPNHF
[6] PFAM. (2014). Summary: PHD finger. Retrieved March 5, 2014 from http://pfam.sanger.ac.uk/family/PF00628.24
[7] PFAM. (2014). Summary: SET domain. Retrieved March 5, 2014 from http://pfam.sanger.ac.uk/family/PF00856.23
[8] The Pawson Lab. (2014). RING domain. Retrieved March 6, 2014 from http://pawsonlab.mshri.on.ca/index.php?option=com_content&task=view&Itemid=64&id=176



