This page was published for Genetics 564 at the University of Wisconsin- Madison
What is homology?
Homology is noted as the structural similarity between traits in different species based on divergence from a common ancestor. Similarities of this nature can be utilized in further study of a particular organism or anatomical area of interest [1][2]. This report focuses on protein homology in particular. Protein homology refers to proteins that show amino acid sequence similarities based on this notion of a common ancestor. These sequence similarities do not necessarily mean functional similarities, however it could provide valuable insight to the function. Functionally equivalent proteins are noted as orthologs. It is key to realize that not all homologs are orthologs, but all orthologs are homologs [3].
Determining protein homology
To search for homologs (the individual species that share protein sequences) from NSD1, Homologene was used to first compare the protein sequence of NSD1(histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific isoform b) with a number of noteworthy model organisms. Comparisons are evaluated on degree of similarity and sorted in order of most similar to least similar [4]. These matches were then tested for validity using the Basic Local Alignment Search Tool (BLAST), an online tool that compares nucleotide sequences to the database of genomes from a plethora of different organisms [5]. A particular species was considered valid if the top hit on a reciprocal BLAST (that is, a search done for the human protein to the species of interest and then back to humans) was the original protein in question. A variety of species were chosen from the results that exhibited a high percent of similarity (approximately >45%) in sequence and a significantly low Expect (E) value. E values close or exactly zero signify a high probability that the sequences chosen out of the database were not purely there by chance and the similarity is actually significant [6].
One notable feature of the protein homologs is the notion of conserved domains. For this sequence in particular, there are a few domains that are seen throughout almost every single homolog, suggesting evolutionary and functional significance.
- PWWP: Pro-tryp-tryp-pro DNA binding proteins that function as transcriptional factors.
- SET: Protein lysine methyltransferase enzymes that help to mediate protein-protein interaction.
- PHD: domains that functions as a zinc-finger involved in mediating protein-protein interactions.
Analysis
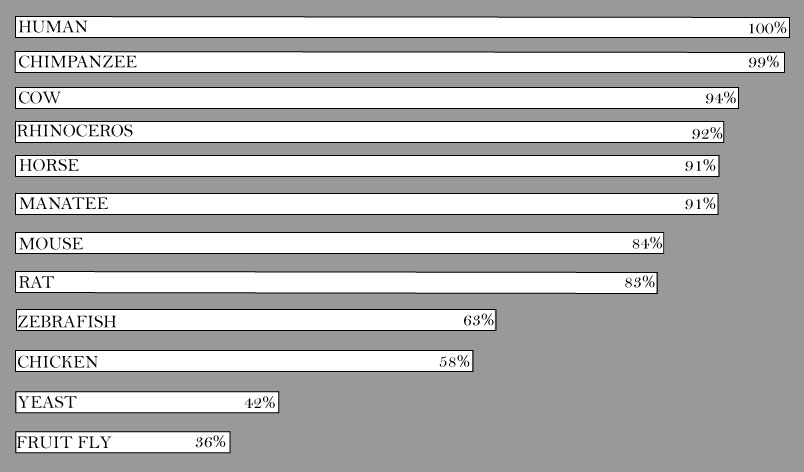
The protein that corresponds with NSD1 is histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific isoform b, better known as NSD1. After running reciprocal BLASTs and analyzing congruence in Homologene, it appears that yet again, humans and chimpanzees see the greatest percent identity at 99% (Figure 1). Given the close relationship of the two species, this seems likely. Larger mammals (rhinoceros, horse, manatee, and cow) continue to show a higher level of homology with that of the human protein. Given that the protein is known to function as a transcriptional regulator and most likely has some associations to development, it seems probable that large mammals share a high percentage of homology. Interestingly, in the gene sequence homology, chickens had a higher percent identity than zebrafish whereas in the protein screen, chickens are the least similar at a low 58% identity. Again, there is a chance that this result could be owing to developmental differences.
Figure 1: The comparison in percent identity as determined by BLAST. Higher congruence in higher percentages.
Protein homolog reference numbers
|
Chicken (Gallus gallus): NSD1
Accession Number: XP_414538.4 AA Number: 2671 Percent Identity: 58% FASTA Rhinoceros (Ceratotherium simum simum): NSD1
Accession Number: XP_004428531.1 AA Number: 2701 Percent Identity: 92% FASTA Fruit Fly (Drosophila melanogaster): Mes-4
Accession Number: NP_001263029.1 AA Number: 1423 Percent Identity: 36% FASTA |
Chimpanzee (Pan troglodytes): NSD1
Accession Number: XP_527132.2 AA Number: 2697 Percent Identity: 99% FASTA Rat (Rattus norvegicus): Nsd1
Accession Number: NP_001100807.1 AA Number: 2381 Percent Identity: 83% FASTA Zebrafish (Danio rerio): nsd1b
Accession Number: XP_005173941.1 AA Number: 1872 Percent Identity: 63% FASTA Horse (Equus caballus): NSD1
Accession Number: XP_001502479.1 AA Number: 2700 Percent Identity: 91% FASTA Manatee (Trichechus manatus latirostris): NSD1
Accession Number: XP_004371211.1 AA Number: 2703 Percent Identity: 91% FASTA |
References:
[1] Eisen, J. (1998) Phylogenomics: improving functional predictions for uncharacterized genes by evolutionary analysis. Genome Res, 8, 163-167. doi: 10.1101/gr.8.3.163
[2] Evolution for Teaching. (2004). The University of Waikato. Retrieved February 3, 2014, from http://sci.waikato.ac.nz/evolution/Homology.shtml
[3] Descorps-Declère, S., Lemoine, F., Sculo, Q., Lespinet, O., & Labedan, B. (2008). The multiple facets of homology and their use in comparative genomics to study the evolution of genes, genomes, and species. Biochimie, 90(4), 595-608. doi: http://dx.doi.org/10.1016/j.biochi.2007.09.010
[4] Homologene Build Procedure. National Center for Biotechnology Information. Retrieved February 5, 2014, from http://www.ncbi.nlm.nih.gov/homologene/build-procedure/
[5] Madden, T., (2011). BLAST Help manual overview. In BLAST Help Manual. Retrieved from http://www.ncbi.nlm.nih.gov/books/NBK52636/
[6] BLAST FAQs. National Center for Biotechnology Information. Retrieved February 5, 2014, from http://www.ncbi.nlm.nih.gov/blast/Blast.cgi?CMD=Web&PAGE_&DOC_#expect
[7] Homologene results. National Center for Biotechnology Information. Retrieved February 15, 2014, from http://www.ncbi.nlm.nih.gov/homologene/?term=NP_071900.2
[1] Eisen, J. (1998) Phylogenomics: improving functional predictions for uncharacterized genes by evolutionary analysis. Genome Res, 8, 163-167. doi: 10.1101/gr.8.3.163
[2] Evolution for Teaching. (2004). The University of Waikato. Retrieved February 3, 2014, from http://sci.waikato.ac.nz/evolution/Homology.shtml
[3] Descorps-Declère, S., Lemoine, F., Sculo, Q., Lespinet, O., & Labedan, B. (2008). The multiple facets of homology and their use in comparative genomics to study the evolution of genes, genomes, and species. Biochimie, 90(4), 595-608. doi: http://dx.doi.org/10.1016/j.biochi.2007.09.010
[4] Homologene Build Procedure. National Center for Biotechnology Information. Retrieved February 5, 2014, from http://www.ncbi.nlm.nih.gov/homologene/build-procedure/
[5] Madden, T., (2011). BLAST Help manual overview. In BLAST Help Manual. Retrieved from http://www.ncbi.nlm.nih.gov/books/NBK52636/
[6] BLAST FAQs. National Center for Biotechnology Information. Retrieved February 5, 2014, from http://www.ncbi.nlm.nih.gov/blast/Blast.cgi?CMD=Web&PAGE_&DOC_#expect
[7] Homologene results. National Center for Biotechnology Information. Retrieved February 15, 2014, from http://www.ncbi.nlm.nih.gov/homologene/?term=NP_071900.2